It's hard to believe we're officially past the half way point of the year! Are you more than half way to achieving your goals? If you're eager to leverage the full potential of your hard earned data and achieve your goals faster, we recommend checking out our top 3 new features that are available now.
Feature 1
Maximise your understanding of protein abundance by leveraging multi-experiment trend analysis - all in one place
With the rapid expansion of both volume and complexity of proteomics data, the ability to easily leverage existing experiments to enhance the interpretation of results is increasingly difficult to do and thus overlooked...until now. Mass Dynamics now enables simultaneous interrogation of multiple experiments, via our Protein Tracer Plot module creating key benefits of:
- Leveraging your existing experiments as a readily searchable internal knowledge base.
- Being able to upload pre-existing results (e.g. from PRIDE) to interrogate your experiment results against external knowledge to see how they compare.
- The ability to interrogate proteins of interest (either stand-alone or as a group) to deduce differential abundance behaviour across experiments.
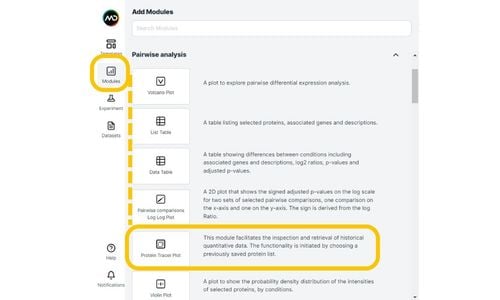
Combined with our current interactive visualisation and collaboration and sharing functionality, your proteomic studies will reach a whole new level of knowledge generation and insights.
----------------
Feature 2
Advance your knowledge of biological processes and systems using our STRING-DB integration to interrogate Protein-Protein Interactions.
We understand one of the traditional key challenges of exploring external knowledge in the context of your experiment typically requires fragmented processes involving importing and exporting data across multiple tools and platforms. That's why we built this seamless integration that enables interactive exploration of protein-protein interactions through a network graph generated via a call to STRING-DB.
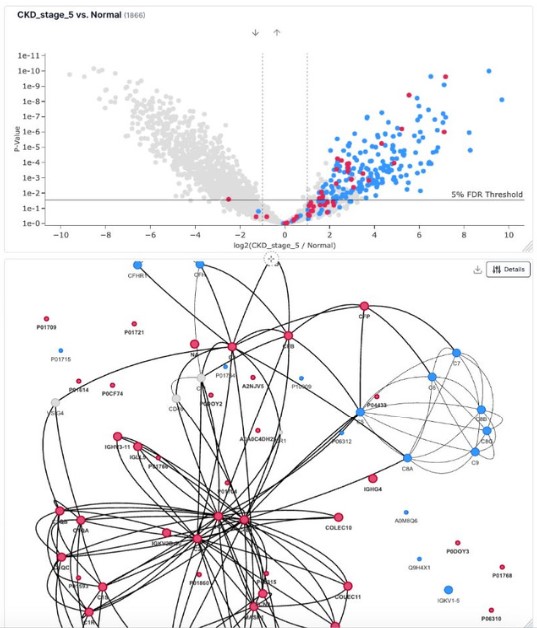
Mass Dynamics empowers scientists by offering a seamless approach, integrating data, conducting advanced network analysis, and delivering premium visualizations, ensuring superior efficiency and reproducibility in research.
----------------
Feature 3
Improve the way you interrogate proteins of interest, with more flexible incorporation of protein lists.
Like an explorer with a well-prepared treasure hunt list, biologists can have lists consisting of proteins known to be associated with specific pathways or functions. These lists can be useful to explore how proteins behave in different conditions, help shed light on understanding the changes that occur when things go wrong, and explore new frontiers in biology.
Creating an unlimited number of protein lists to interrogate across your experiments in Mass Dynamics is now even easier. This includes the ability to:
- Use one Protein List in different analysis modules
- Use multiple lists to track protein behaviour across different analysis modules
- Import protein lists from experiments performed inside or outside of Mass Dynamics
- Allow your collaborators to see the same protein selections when you share an experiment
Watch Head of Product Design, Brad Green describe Protein Lists as one of the fundamentals of using Mass Dynamics Discovery Service (Timestamp 1:21).
 What's New? Q2-23">
What's New? Q2-23">