It wasn’t a trade show or a marketing showcase. The room was intimate - directors, VPs, heads of platform, chemists, bioinformaticians. The kinds of people who carry decisions that ripple across programs.
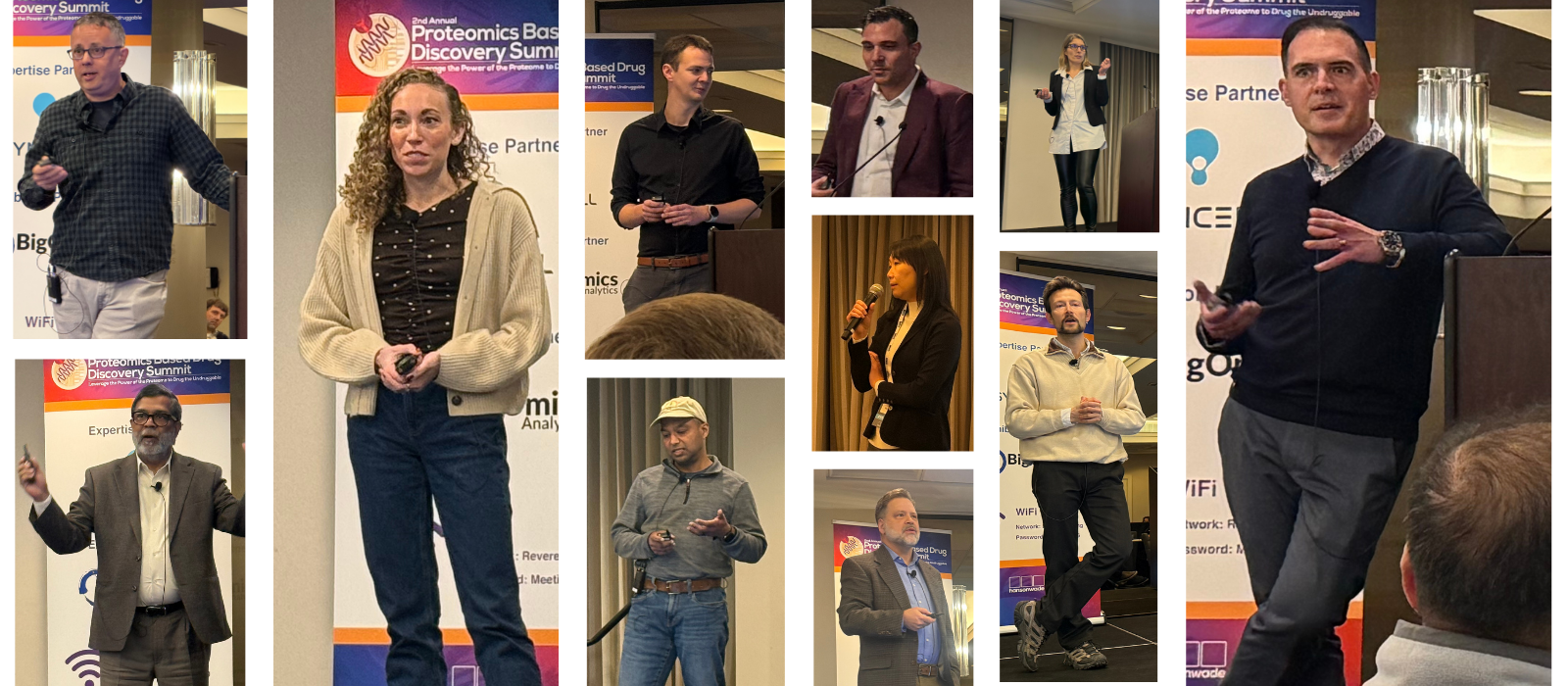

The ambition of the summit was clear: harness next-generation proteomics to move faster and smarter through drug discovery. Balance throughput and sensitivity. Automate sample processing. Improve PTM analysis. Define gold-standard methodologies. Expand the druggable proteome.
And on paper, proteomics has never looked stronger.
Novartis - ProFound: $25M upfront/near-term, with up to $750M per target in downstream milestones. Illumina - SomaLogic: up to $425M including milestones. Whatever you think of the individual bets, to me the signal is unmistakable:
Proteomics has moved from exploratory to strategic.
Measurement has matured. Throughput has improved. Sensitivity is no longer the limiting factor it once was.
But sitting in that room, listening across very different talks, I kept coming back to the same thought:
Once a field matures, the constraint shifts. For years, the bottleneck was measurement - could we go deeper, faster, more sensitively?
What became clear to me in Boston is that the bottleneck has moved.
-
Throughput is now outpacing our ability to interpret and reuse data.
-
Discordance across peptides, platforms, and teams erodes confidence.
-
Experimental context isn’t consistently captured or carried forward.
-
Data analysis capacity hasn’t scaled at the same rate as data generation.
The next breakthrough won’t come from deeper spectra alone. It will come from infrastructure that makes insight durable, comparable, and decision-ready.
That shift, from measurement to coordination, is why I attended.
With AI advancing rapidly and new computational tooling appearing almost monthly, expectations around decision velocity inside pharma are increasing. At the same time, capital discipline is tightening and timelines are compressing. The pressure to convert data into confident decisions - quickly - has never been higher.
When scientific maturity, capital commitment, technological acceleration, and enterprise pressure converge like this, the bottleneck moves.
In Boston, I felt that shift happening in real time.
I wanted to document what was being said in that room — at this inflection point. Not the marketing narrative, or the hype cycle. But the underlying tension of what’s working, where friction remains, and what we, collectively, need to address if proteomics is going to move from powerful to indispensable.
Patients are waiting, and the pace at which we turn biological insight into therapeutic impact matters.
By the end of the day, one thing felt clear to me:
The hard problem in proteomics is no longer measurement - it’s coordination.

Setting the tone: decision-making under real constraints
An Chi (Merck) opened the day as Session Chair and, in many ways, quietly set the frame for everything that followed.
She didn’t lean on urgency or crisis, nor did she oversell proteomics as a silver bullet.. Instead, she grounded the conversation in something more demanding: decision-making under real constraints. She encouraged discussion, shared learnings and debate on:
-
Where has proteomics genuinely influenced programs? Or where has it under-delivered?
-
And what does it take for data to move from technically impressive to operationally trusted?
Her emphasis wasn’t on capability, but on accountability. She encouraged everyone to share what works, and admit what doesn’t, and to be honest about tooling and workflow gaps rather than hiding behind performance metrics.
The tone was steady, pragmatic, and quietly optimistic. And that mattered because the rest of the day reinforced her central point: the measurement layer has matured - but our ability to learn collectively hasn’t matured at the same pace.
Making proteomics first-class inside a global pharma
One senior pharma leader’s talk was repeatedly described by others in the room as inspiring. Not because of flashy science necessarily, but because of what had been achieved organizationally: making proteomics a first-class citizen inside a therapeutic pipeline.
A few things stood out:
-
Throughput is no longer the bottleneck (100–300 samples/day is now feasible).
-
The job is not to deliver data - it is to deliver insight.
-
Data matrices shouldn’t be “thrown over the fence.”
-
Black-box thinking is the wrong model for proteomics.
There was a quiet maturity in that framing.
For years, the field has focused on whether we could generate data at scale - more depth, more samples, more sensitivity. At one point, a well-known line from Jurassic Park was invoked:
“Scientists were so preoccupied with whether they could, they didn’t stop to think if they should.”
The point wasn’t to argue against capability. It was to argue for responsibility.
Automation without alignment doesn’t create progress. It accelerates output - and without shared interpretation, that output becomes noise.
The emphasis for this same pharma talk was on flow: movement of data across teams, and systems that reduce friction between scientists rather than replacing them. The subtext was clear: automation without coordination accelerates confusion.
What made this resonate was that it wasn’t theoretical. Teams operating at this scale have generated petabytes of data and lived the coordination pain firsthand.
What I took from this pharma-focussed talk was simple: at scale, insight doesn’t just require throughput — it requires structure.
The theme of “scale demanding structure” surfaced again throughout the day.

Immunopeptidomics, scale, and the coordination challenge
Immatics operates at the sharp end of immunopeptidomics - mapping HLA-presented peptides to identify targets for T-cell–based immunotherapies. It’s a domain where precision matters, false positives are unacceptable, and biological interpretation is inseparable from the data itself.
Christoph Schräder's (Immatics) talk was a reminder of just how specialized (and demanding) this work is.
A few sharp signals:
-
Absolute quantification is run as a default, not an exception.
-
Data volumes already exceed what can realistically be surfaced in talks or figures.
-
They operate at a scale where around 20 bioinformaticians support the platform.
That number stuck with me as a signal of how much intellectual labour is required to keep such a system functioning. There’s no question those bioinformaticians are generating insight. In fact, their expertise is what makes immunopeptidomics viable at this level at all. But when a field becomes this complex, a meaningful portion of expert time inevitably goes into coordination work:
-
maintaining bespoke pipelines
-
reconciling edge cases
-
preserving methodological nuance
-
translating results across teams and programs
All necessary. All valuable. And all largely invisible.
The deeper question this raises for the field isn’t about people - it’s about structure:
As complexity grows, how much expert effort is spent advancing biology, and how much is spent holding the system together?
Keeping methods in-house, as Immatics does, is a rational and defensible choice, especially when IP sensitivity and biological subtlety are high. And it also means that scaling insight often scales people, rather than reducing the coordination burden around them.
Christoph didn’t frame this as a problem, nor did he need to. The signal was clear for us to see: as immunopeptidomics matures, the constraint isn’t data generation or talent. It’s how well we support experts so their time is spent on biology.
.png)
Proof, skepticism, the biology
Aethon Therapeutics sits at the intersection of chemistry, biology, and systems rigor - focused on building drugs where mechanism has to be proven, not assumed. Their work forces uncomfortable questions early: did the target actually make it to the cell surface, did the degrader do what we think it did, and can we demonstrate that with evidence rather than inference?
Lauren Stopfer (Aethon Therapeutics) spoke candidly about early skepticism - the need for direct proof that what they believed was happening biologically was actually real. To answer that, they invested deeply in orthogonal validation, including SPR and mass spectrometry, to show targets truly reached the cell surface.
Importantly, while the FDA-approved drug in this class is not theirs, it demonstrates that this therapeutic approach has real commercial and clinical viability. Others in the space (including Aethon’s efforts) are aimed at unlocking entirely new classes.
Two moments stood out from Lauren's presentation:
- Investors need proof, not just narratives.
- In Q&A, she said:
I wish more companies would share what their degraders actually look like.
That comment landed hard. Because it wasn’t about IP bravado - it was about collective learning velocity. If we don’t share structures, signatures, and failures, everyone pays the coordination tax alone.
.png)
Iteration at scale and executive pressure
Uthpala Seneviratne (AstraZeneca) has been involved in two FDA-approved drugs, and his perspective reflected that maturity. They run DMTA cycles repeatedly - targeted, high-throughput approaches designed to get to leads efficiently.
Discovery still happens, but it’s deliberately bounded. Off-targets and safety are handled elsewhere in the pipeline.
What came through clearly in Uthpala's talk:
-
Executives care deeply about cost and efficiency.
-
Proteomics is trusted, but only when it supports decisive iteration.
-
Targeted approaches are about control, not curiosity.
This wasn’t anti-discovery. It was a reflection of pressure: once scale and stakes increase, coordination becomes more valuable than optionality.
.png)
When depth, discordance, and truth collide
Satya Saxena's (Eisai) talk was one of the most technically provocative of the day.
Working in neurology - where you can’t biopsy brains - forces uncomfortable trade-offs. He spoke about combining platforms to extend proteome depth, about studies where overlap between methods was surprisingly low, and about the danger of filtering out “outlier” proteins too early.
He used a word I hadn’t heard much in this context before: discordant.
-
Discordant peptides.
-
Discordant signals.
-
Discordant interpretations.
His call was clear: we need to look more at peptide-level data, not just protein summaries. And we need to be honest about where confidence actually comes from.
.png)
From invention to industrialization: when proteomics becomes decision-critical
Arvinas is the original standard-bearer of targeted protein degradation. That maturity changes the role proteomics plays.
John Corradi's (Arvinas) talk made one thing unambiguous: at Arvinas, proteomics is no longer a supporting actor - it’s a primary efficacy readout.
In TPD, this is existential. The biology lives or dies on confidence:
-
Is the target degraded?
-
How selectively?
-
In which context?
-
With what downstream consequence?
John walked through how proteomics is used across the arc, from lead optimization through translational and clinical work, alongside complementary platforms such as SomaScan. What stood out wasn’t methodological novelty; it was the weight placed on interpretation.
When proteomics becomes decision-critical, familiar technical debates take on organizational change. Disagreements over thresholds or discordant peptides require decisions that must remain defensible months or years later, across teams and programs.
At that scale, continuity of understanding matters more than novelty. Not more data. Not flashier methods. But coordination, so that insight doesn’t fracture as pipelines grow.
.png)
Data analysis is now the bottleneck
Regeneron operates at significant therapeutic breadth - oncology, infectious disease, autoimmunity, metabolism - and Robert Salzler's (Regeneron) talk reflected that operational maturity. He didn’t need to define immunopeptidomics since the audience already understood the landscape from previous presentations.
What he did emphasize was stark:
-
Automation and mass spectrometry throughput are largely solved
-
Data analysis is now the dominant bottleneck
That shift matters. When instrumentation becomes reliable and throughput increases, attention inevitably moves downstream - interpretation, integration, and communicating statistical confidence.
Robert pointed to repositories, genomic integrations, ribo-seq overlays, and persistent FDR challenges. Especially when proteomics is layered with genomic data or custom database searches, search space expands, thresholds tighten, and confidence becomes harder to convey.
The message aligned with much of what had been building throughout the day: the field has matured technically, but the systems that help teams interpret, reconcile, and reuse data have not scaled at the same pace.
It’s no longer about generating spectra. It’s about converting complex, multi-omic outputs into durable, shared understanding.
A scientific inflection point
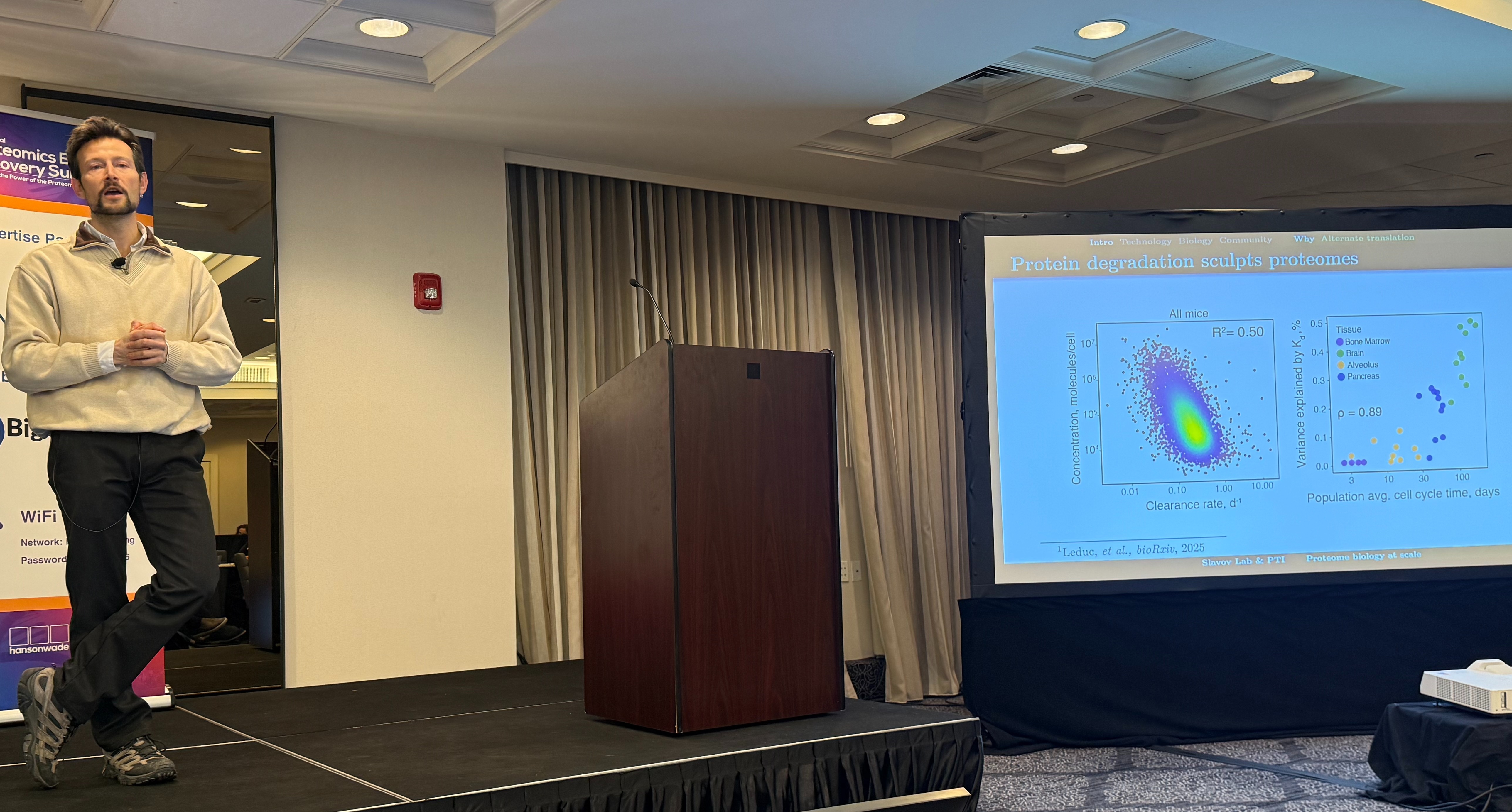
Nikolai Slavov’s (Parallel Squared Technology Institute) talk marked a scientific inflection point in the field.
For years, single-cell proteomics was described as “almost there”, limited by sensitivity, throughput, or hardware. What Slavov showed is that many of those constraints have shifted. Through real-time instrument control, parallelisation strategies like plexDIA, and engineered mass tags, single-cell proteomes can now be measured at meaningful scale, even on legacy instruments.
But the deeper shift doesn’t appear to be technical, it appears to be systemic.
As throughput increases, the constraint moves. When measurement is no longer the primary barrier, the challenge becomes interpretation: prioritising signals, resolving discordant peptides, integrating turnover and PTMs, and preserving context across perturbations and time.
One example captured the biological weight of this shift. By combining single-cell proteomics with metabolic labelling, his work suggests that in long-lived cells like neurons, protein abundance is often governed by degradation rather than transcription.
“My neurons were with me when I was born, and they’ll be with me when I die.”
It’s a reminder that turnover biology is foundational, not theoretical.
Slavov was also clear-eyed about adoption. While the science is advancing rapidly: cost, operational complexity, and standardization remain real barriers to broad implementation. The next phase isn’t about proving that single-cell proteomics works. It’s about building systems that make it scalable and decision-ready.

Expanding resolution: from single-cell to nanoscopic
If Slavov showed that single-cell proteomics is crossing a threshold, Syncell pushed the conversation even further - into the nanoscopic.
Daniel Dlugolenski's (Syncell) framing was clear: “Measuring biology at the scale where mechanisms actually occur”, and their focus on surfaceome technology stood out:
-
60–70% of FDA-approved drugs target surface proteins.
-
Yet surface proteins account for only ~2% of total protein abundance.
-
And roughly one-third of the human proteome encodes surface proteins.
Daniel showed high-precision biotinylation of the immune synapse, isolating proteomes at the interface between interacting cells.
That level of resolution can open new discovery territory, and also increases complexity. The open question is whether coordination can accelerate at the same pace as this technology.
So what was the real takeaway for me?
Across pharma, biotech, platform builders, immunologists, and single-cell pioneers, one pattern kept emerging.
Proteomics is no longer trying to prove itself. It works, it scales, and it informs real therapeutic decisions.
And once that happens, the constraint moves. The friction doesn't appear to be in the instruments anymore. It’s in how insight is carried forward.
Throughout the day, that tension surfaced in different language, but it pointed to the same underlying issues:
- Critical experimental context isn’t consistently captured.
- Interpretation shifts across teams and time.
- Integrating proteomics with genomics and other modalities multiplies complexity.
As data continues to be generated at unprecedented volumes:
-
Depositing it somewhere doesn’t make it usable.
-
A shared file format doesn’t create shared understanding.
-
Responsibility for making data truly reusable often falls between teams.
I believe the gap isn’t technical, it’s structural.
None of this was framed as crisis. It felt more like the growing pain of a field moving from innovation to infrastructure. That shift matters because when proteomics becomes strategic (through informing portfolio decisions, clinical direction, and capital allocation), coordination stops being optional and becomes foundational.
Other fields have crossed this chasm before. The day included discussions about how genomics didn’t become indispensable because sequencing got cheaper. Instead, it became indispensable because shared reference frameworks and systems made complexity navigable at scale.
Proteomics seems to be at that moment now.
What gave me confidence in Boston wasn’t a single platform or method. It was the group's alignment:
- A belief that proteomics belongs at the decision table.
- A shared frustration with fragmentation and re-work.
- A desire to learn together, not just publish alone.
I believe what matters now is acknowledging where the real work sits — and being specific about what coordination actually requires. The future of proteomics will be shaped less by how fast we measure and more by how intentionally we build the infrastructure that allows insight to survive across disciplines, data layers, and decision-makers.
If coordination is now the constraint, then we need to be explicit about what that implies.
Not more plots. Not more isolated workflows. Not more dashboards layered on top of fragmentation.
We need three things.
1. Context capture.
Not just files and outputs, but experimental decisions and analytical reasoning. Insight decays when context is lost.
2. Reproducible interpretation.
Versioned thresholds, peptide-level traceability, and clear provenance from signal to biological claim. Confidence must survive scrutiny and time.
3. Cross-team continuity.
Infrastructure that allows insight to compound - across programs, across tooling shifts, and across organisational change.
None of this is glamorous. All of it is infrastructural - and infrastructure is what turns a powerful technology into a dependable one.
Proteomics has already proven it can measure. Now it needs to prove it can coordinate.
I’m planning to follow and share what we learn as this plays out through 2026. Not because the answers are obvious, but because the questions are now the right ones.
---------
This blog post was produced by Paula Burton using a combination of notes from presentations, discussions and insights. Final compilation was completed with assistance from ChatGPT 5.2. Each speaker reviewed their own section. No speaker reviewed the full piece end-to-end, and all synthesis and takeaways are my own. Any errors or omissions are unintentional, and the content is provided for informational purposes only. The views, thoughts, and opinions expressed in this text belong solely to the author, and not necessarily to the author's employers, organization, committees or other group or individual